Debugging nodes¶
Debugging nodes using PyCharm¶
PyCharm is an external IDE (not included in Sympathy) which provides good debugging support for Sympathy.
In order to debug Sympathy in PyCharm follow these steps.
Create a new Project
In the New Project dialog: configure the Python interpreter (python.exe) used to run Sympathy as the Project Interpreter.
Start by checking Previously configured interpreter. Then click Add interpreter -> Add Local interpreter -> System Interpreter and finally browse to and choose the interpreter.
You may have to wait some time for indexing to complete before the interpreter is ready to use.
The interpreter can also be changed afterwards from Settings -> Project -> Project Interpreter.
In Settings -> Project -> Project Structure, add the folders containing the code that you want to debug using Add Content Root. For example, in order to debug anything from Sympathy itself (including the standard library nodes) add <Python>/Lib/site-packages/sympathy and <Python>/Lib/site-packages/sylib. Then you may have to wait for another indexing.
In Run->Edit Configurations, create a new configuration that uses the Project Interpreter. Choose a name that is easy to remember. Use Module name instead of Script path to specify the target. Module name: sympathy, and Parameters: gui. Then uncheck Add content roots to PYTHONPATH to avoid module and package name conflicts. The same goes for Add source roots to PYTHONPATH should it give rise to conflicts. Generally it is important to be careful when adding to PYTHONPATH since it can result in problems in third-party code. Finally choose an appropriate working directory. The configuration is now complete!
In Run, choose Debug <your-configuration-name>, this will start Sympathy in debug mode. In PyCharm, you can set breakpoints in files by clicking on the left margin, see https://www.jetbrains.com/help/pycharm/using-breakpoints.html#set-breakpoints. When execution reaches a line with a breakpoint, PyCharm will stop there and give you control.
For example, to debug the execute part of the Hello world example node, open the node’s file, <Python>/Lib/site-packages/sylib/nodes/sympathy/examples/node_examples.py in PyCharm, locate the execute method and on the following line, containing: print(‘Hello world!’) insert a breakpoint. In Sympathy, create a Hello world example node and execute it. PyCharm should eventually stop at the line with the breakpoint.
Debugging nodes using Visual Studio Code¶
Visual Studio Code (VSCode) is an external IDE (not included in Sympathy) which is versatile and can be adapted to support Sympathy debugging. This guide was created using VSCode version 1.61.1. We therefore recommend upgrading to at least this version before attempting this tutorial.
In order to debug Sympathy in VSCode follow these steps:
Install the Python plugin for your operating system here and select the Python Interpreter.
It is assumed that Sympathy is installed via installer, so no additional Python installation is required since Sympathy comes with its own Python distribution. This distribution can be used in VSCode. Then select the correct Python Interpreter by clicking on the bottom-left corner as shown below. Alternatively, press Ctrl+Shift+P and then search for Python: Select Interpreter and find the path to your Python installation.
For more information about Python support in VSCode you can check out: Python support help.
Create a workspace in VS Code.
All that is required to “create” a workspace is to open a folder within VS Code that has not been used for any VS Code projects. Once a folder has been opened, VS Code will automatically keep track of things such as your open files and editor layout so the editor will be as you left it when you reopen that folder. To read more about workspaces in VS Code, you can read through the following: Workspaces.
Create the launch.json file.
Although it is possible to configure debugging globally using settings.json, we recommend configuring Sympathy debugging at the Workspace level. To add Sympathy debugging configuration to your project, we will need to work with a launch.json file in the project’s .vscode folder. Create it by selecting the “Run and Debug” mode on the left-hand side menu in VS Code, and then select “create a launch.json file” as shown in the figure below. You can then follow the steps by selecting the workspace name, the debug configuration (any since this will be replaced) and then launch.json should open with a template. These steps are also illustrated below.
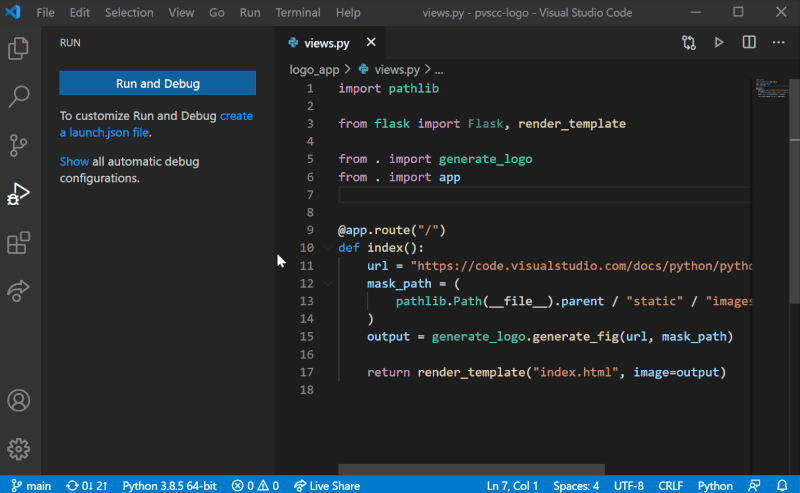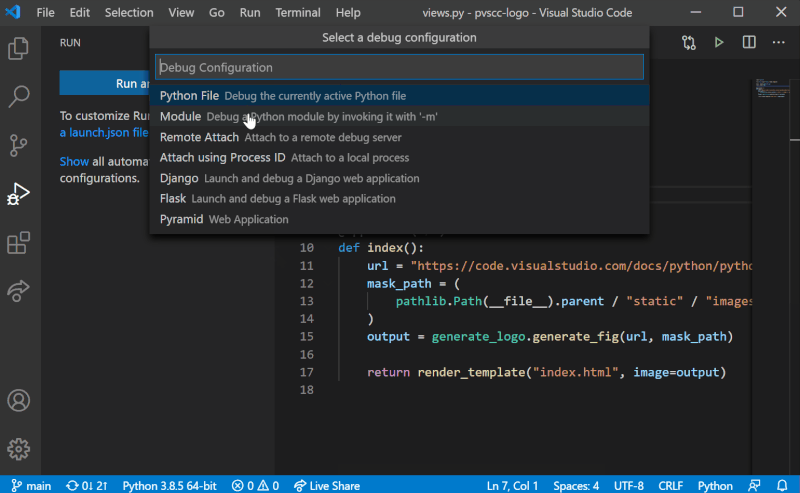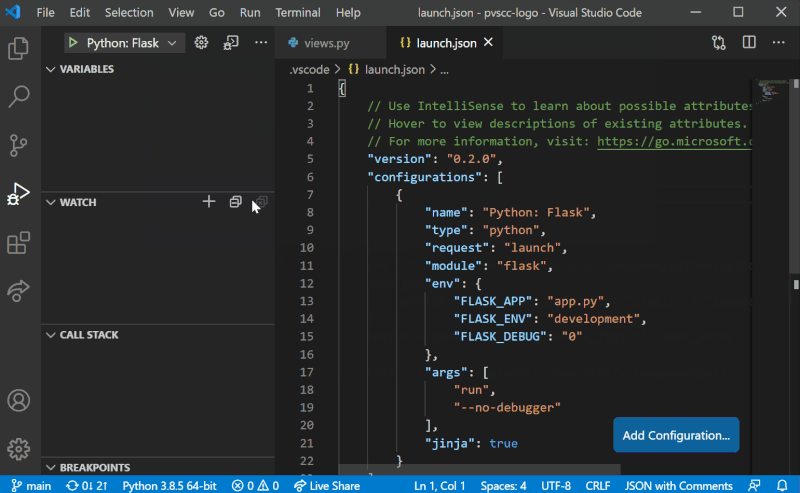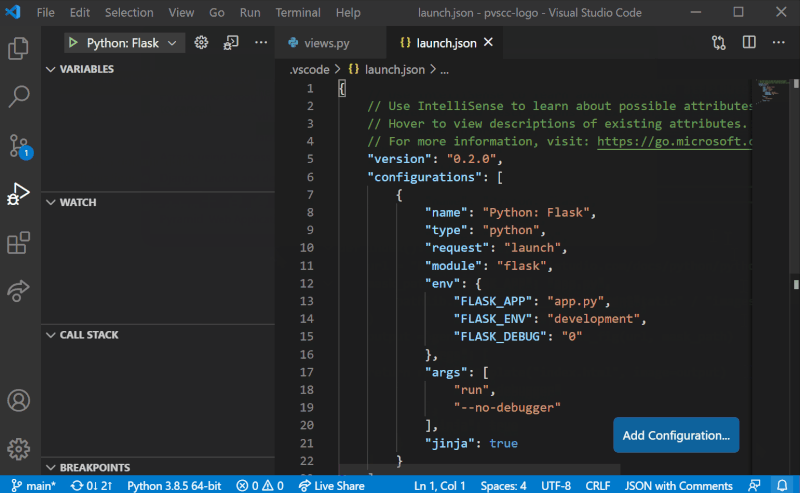
Replace the entire contents of launch.json with the following:
{ "version": "0.2.0", "configurations": [{ "name": "Python: Debug Sympathy", "type": "python", "request": "launch", "module": "sympathy", "python": "<Path to Sympathy for Data>/Python/python.exe", "args": ["gui"], "console": "integratedTerminal", "justMyCode": true, "subProcess": true }] }
Replace <Path to Sympathy for Data> with your respective Sympathy installation directory. Note that default value for “justMyCode” is true, which will skip debugging Sympathy platform code. This means only your own node code will be debugged.
Switch to “Run mode” (Ctrl-Shift-D) in the VSCode IDE, and choose the created debugging configuration in the drop-down list on top. Start the debugging by clicking on the green “Play” symbol:
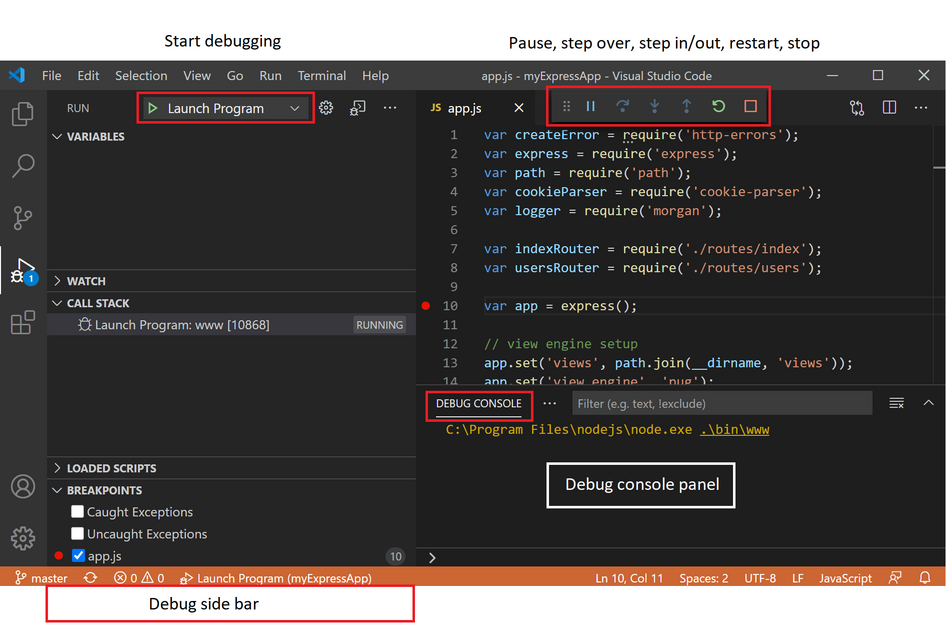
If successful, the Sympathy for Data splash screen should appear after the processes have spawned within VS Code. For more advanced or general information on Python debugging support in VSCode visit: VSCode Debugging Documentation.
Profiling nodes and workflows¶
If your node or workflow is running too slow you can run the profiler on it to see what parts are taking the most time. If you have configured Graphviz, see Configuring Graphviz, you will also get a call graph.
To profile a single node simply right-click on a node that can be executed and choose Advanced->Profile. This will execute the node and any nodes before it that need to be executed, but only the node for which you chose Profile will be included in the profiling. To profile an entire workflow go to the Controls menu and choose Profile flow. This will execute all nodes in the workflow just as via the Execute flow command. After either execution a report of the profiling is presented in the Messages view. Profiling of single subflows is similar to profiling of single nodes but include all the executable nodes in the subflow.
The profile report consists of a part called Profile report files and a part called Profile report summary.
Profile report files¶
The Profile report files part of the profile report consists of two or four file paths. There is always a path to a txt file and a stats file, and also two pdf files if Graphviz is configured, see Configuring Graphviz. The txt file is a more verbose version of the summary but with full path names and without any limit on the number of rows. The first pdf file contains a visualization of the information in the summary, also called a call graph, and the second a call graph of flows and nodes.
The function call graph contains a node for each function that has been called in the profiled code. The color of the node gives you a hint about how much of the total time was spent inside a specific function. Blue nodes represent functions with low total running time and red nodes represent functions with high total running time. The nodes have the following layout:
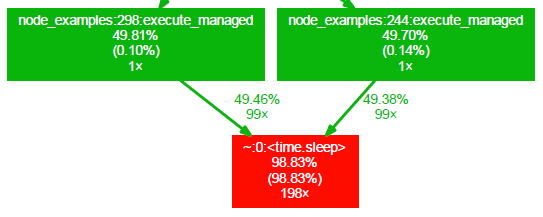
First row is the name of the function. Second row is the percentage of the running time spent in this function and all its children. Third row (in parentheses) is the percentage of the running time spent in this function alone. The forth row is the total number of times this function was called (including recursive calls).
The edges of the graph represent the calls between functions and the label at an edge tells you the percentage of the running time transferred from the children to this parent (if available). The second row of the label tells you the number of calls the parent function called the children.
Please note that the total running time of a function has to exceed a certain cut-off to be added to the call graph. So some very fast workflows can produce almost empty call graphs.
A third file will also always be provided with the file ending “.stats”. This file contains all the statistics that was used to create the summary and the call graph. To begin digging through this file open a Python interpreter and write:
>>> import pstats
>>> s = pstats.Stats('/path/to/file.stats')
>>> s.print_stats()
For more information look at the documentation for the Stats class.
Profile report summary¶
The summary contains a row for each function that has been called in the profiled code. Several calls to the same function are gathered into a single row. The first column tells you the number of times a function has been called. The next four columns measure the time that it took to run a specific function. In the last column you can see what function the row is about. See https://docs.python.org/2/library/profile.html for details on how to interpret this table.
The summary also includes up to 10 node names from nodes included in the profiling and an indication of the number of nodes that were omitted to save space.
Writing tests for your nodes¶
As with any other code, writing tests for your nodes is a good way of assuring that the nodes work and continue to work as you expect.
Test workflows¶
The easiest way to test the execution of your nodes is to add them to a workflow (.syx) and put that workflow in the test flows folder. See Libraries for information about where to put test flows in your library.
In some cases it can be enough to test that the flows can execute without producing exceptions or errors, in other cases, the actual data produced need to be checked. For comparing data, Conditional error/warning and Assert Equal Table may come in handy.
Unit tests¶
It is also a good idea to write unit tests to ensure the quality of your
python modules. Put unit tests in the test folder (see Libraries). If the
tests are named correctly they can be run automatically by installing and
running the Python package pytest. Cd into the library directory and run
pytest to run all unit tests there:
cd my_library
pytest test
See https://docs.pytest.org/en/6.2.x/goodpractices.html#conventions-for-python-test-discovery for more details about how to name your unit tests for pytest to find them automatically.
For example a unit test script that tests the two functions foo() and
bar() in the module boblib.bobutils could be called
test_bobutils.py and look something like this:
import numpy as np
import pytest
import boblib.bobutils
def test_foo():
"""Test bobutils.foo."""
assert boblib.bobutils.foo(1) == 2
assert boblib.bobutils.foo(0) == None
with pytest.raises(ValueError):
boblib.bobutils.foo(-1)
def test_bar():
"""Test bobutils.bar."""
input = np.array([True, False, True])
expected = np.array([False, False, True])
output = boblib.bobutils.bar(input)
assert all(output == expected)
For more help on how to write unit tests take a look at the documentation for
the pytest module at https://docs.pytest.org.
